Professor Kihara and Lab Compete at International Conference
07-13-2005


Professor Daisuke Kihara and his lab fared the best in scored competition for their Protein Function Prediction (PFP) at the International Conference on Intelligent Systems for Molecular Biology in Detroit, Michigan. Their work beat notable entries from David Eisenberg's lab at UCLA, Andrej Sali's lab at UCSF, Adam Godzik's lab at the Burnham Institute, and the European Bioinformatics Institute at EMBL. PFP is a web server designed to give sequence and structural information for a series of proteins. The results are listed as the top ten most probable Gene Ontology annotations in the Biological Process, Molecular Function, and Cellular Component categories.
In the age of genome projects and high-throughput proteomics experiments, biologists are turning more and more to bioinformatics to help organize data and guide experiments. Function prediction of identified but yet uncharacterized proteins is a key element in this relationship, providing what some refer to as "computationally-assisted hypothesis generation."
A special interest group meeting on Automated Function Prediction at the recent International Conference on Intelligent Systems for Molecular Biology was held June 24-29. The conference hosted an assessment of web servers that annotate a query protein sequence or structure. The format of the assessment closely resembled that of the widely-known CASP (Critical Assessment of Techniques for Protein Structure Prediction), but was the first of its type. Participants were required to provide a fully automatic server to which the organizer submitted sequence and structural information for a series of proteins for which functions were known but not yet published. The accuracy of each prediction was assessed by calculating the degree of specificity the predicted annotation shared with the known function annotation in the Gene Ontologies, a categorized, hierarchical vocabulary of biological function terms.
PFP uses PSI-BLAST, the traditional tool used by biologists for inference of homology, and FAMs (Function Association Matrices), a novel functional data mining tool, to produce a unique score for each possible annotation, providing both accuracy and strong coverage of functional space. In addition to participating in the assessment, Kihara and graduate student Troy Hawkins gave an invited talk, "The use of context-based functional association in automated protein function prediction methods", and presented two posters describing their work.
Dr. Daisuke Kihara runs the Purdue Bioinformatics Laboratory. He is an Assistant Professor in Biological Sciences and Computer Science. Troy Hawkins is a second year PhD student in Biological Sciences. The PFP server is online at http://kiharalab.org/web/pfp.php.
Congratulations to the Purdue Bioinformatics Lab!
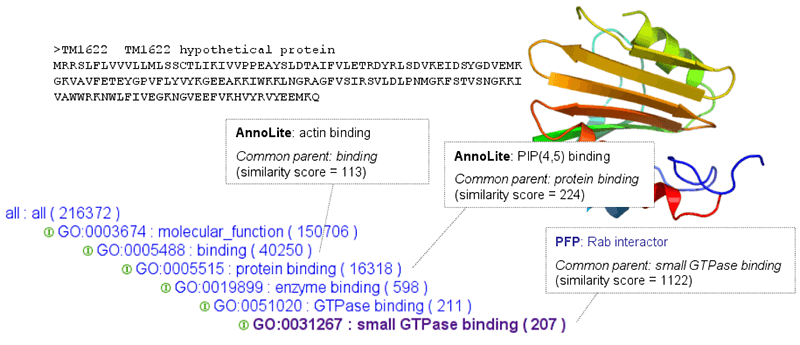
Figure: The prediction accuracy of each participating server was calculated by a similarity score of the predicted annotation to the accepted annotation based on the Gene Ontology (GO) tree. Shown here is the assessment of one of the target proteins, TM1622. PFP predicts a very specific molecular function annotation for this target protein, a GTP binding regulator, while other servers predict more general annotations or no annotations at all.