CS Grad Student, Professors Earn Best Paper Award at Prestigious ACM Bioinformatics Conference
10-15-2015
Writer(s): Jesica E. Hollinger
Asish Ghoshal, a graduate student in Computer Science and Professors Ananth Grama (Computer Science), Saurabh Bagchi (Electrical and Computer Engineering), and Somali Chaterji (Computer Science) took top honors with their paper “An Ensemble SVM Model for the Accurate Prediction of Non-Canonical MicroRNA Targets” at the 2015 ACM Conference on Bioinformatics, Computational Biology, and Health Informatics (ACM-BCB) held in Atlanta, September 9-12.
The ACM-BCB conference offers a forum for premier interdisciplinary research linking computer science, mathematics, statistics, biology, bioinformatics, and health informatics. Additionally, the conference showcases leading-edge research on new technologies and techniques for gathering, processing, analyzing, and modeling these big data, from bench to bedside, for a variety of scientific, clinical, and healthcare applications.
Dr. Chaterji described the team’s research and their creation of “Avishkar”, a machine-learning, support vector machine-based model, to predict both canonical and non-canonical (non-conventional) miRNA-mRNA interactions. The word Avishkar comes from the Sanskrit word, meaning discovery.
“Avishkar extracts features from high-throughput biological databases that harbor RNA regulatory interactions and uses them to predict which RNAs will affect gene expression and to what extent,” Chaterji said. “We used non-linear support vector machines (SVMs) in order to minimize bias and scale Avishkar up to the large biological data sizes through a biologically-driven parallelization strategy. Through this, we achieve the best-in-class recall (true positive rate), with an improvement of over 150%, over the best competitive protocol,” she added.
MicroRNAs, shortened as miRNAs, are short 20-24 nucleotide, endogenous regulatory RNAs that modulate gene expression and through that, play a role in a host of diseases, both simple (involving a single gene) and complex (e.g., type II diabetes, Alzheimer’s, cardiovascular). Chaterji said, “The deficiency of current computational target-prediction approaches has been a bottleneck in mapping out the full functional repertoire of the RNA molecules. Thus, via deciphering which miRNAs will target which genes, it is possible to develop therapeutic treatments. So, our algorithm is key to developing novel miRNA-based therapeutics, some of which are already being pursued as clinical candidates for cardiac repair and treating atherosclerosis,” she added.
She explained that all cells in the body have the same set of genes encoded in their DNA, but it is the presence of these cell-type specific post-transcriptional regulators, miRNAs, further encoded in the DNA, that fine-tune gene expression in more than the 200 different cell types in the human body.
Professor Chaterji works collaboratively with Professor Ananth Grama to use her biomedical engineering and genomics data mining skills to solve problems in computational genomics.
This Nature Review web site demonstrates the biological mechanisms underlying miRNA action: http://www.nature.com/nrg/multimedia/rnai/animation/index.html
Their winning paper can be read at: https://www.cs.purdue.edu/homes/schaterj/papers/avishkar_bcb15.pdf
Interested readers can access their code through their open-source software release: https://bitbucket.org/cellsandmachines/kernelsvmspark
A part of the team can be viewed at: https://www.cs.purdue.edu/homes/schaterj/pics/acm_bcb_award_2015.jpg
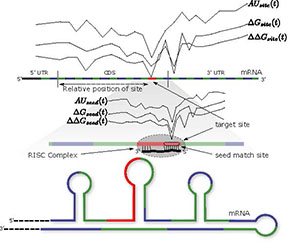
* Figure 1 as seen above: Sequence and Thermodynamic Curves Capturing the Logistics of MicroRNA-Messenger RNA Interactions (courtesy: Ghoshal et al., 2015), as conceptualized by Avishkar, in https://www.cs.purdue.edu/homes/schaterj/papers/avishkar_bcb15.pdf